LabRoots has grown into the world’s largest series of virtual events within the Life Sciences and Clinical Diagnostics community.Take advantage of Eclipse Bio’s unique, best-in-class RNA genomic technologies to optimize your biological and therapeutics research. Founded in 2008, LabRoots emphasizes digital innovation in scientific collaboration and learning, and is a primary source for current scientific news, webinars, virtual conferences, and more. Contributing to the advancement of science through content sharing capabilities, LabRoots is a powerful advocate in amplifying global networks and communities. LabRoots is the leading scientific social networking website, which provides daily scientific trending news and science-themed apparel, as well as produces educational virtual events and webinars, on the latest discoveries and advancements in science. Lexogen is based in Vienna, Austria, and has a subsidiary in Greenland, NH, US. Other products include SPLIT RNA Extraction Kit, TeloPrime Full-Length cDNA Amplification Kit, RiboCop rRNA Depletion Kit, Spike-in RNA Variant Control Mixes (SIRVs), and Mix2 RNA-Seq Data Analysis Software for highly accurate concentration estimates of gene isoforms. All library preparation protocols are based on Lexogen’s proprietary technologies which eliminate the need for RNA or cDNA fragmentation, allowing to preserve superior strand-specificity (>99.9%) and enabling exceptionally rapid turnaround times. QuantSeq library preparation kits are for 3’ mRNA-Seq and target-specific sequencing. SENSE kits are for whole transcriptome sequencing and are suitable for anaylsis of mRNA and total RNA from intact RNA samples, as well as highly degraded and FFPE samples. Lexogen offers two types of sequencing library preparation kits. Its portfolio includes innovative molecular biology kits, software, and services for RNA-Seq. Lexogen is a transcriptomics and Next Generation Sequencing (NGS) company, focusing on the development of technologies for complete transcriptome sequencing. To read more about this event, learn of the continuing education credits offered, or to register for free, click here. LabRoots will host the event on June 28, 2017, beginning 9:00 a.m. He is collaborating with several research groups performing data analysis and developing, an interactive research platform for the study of post-transcriptional modifications. Rot obtained a Masters in computer science from the University of Ljubljana, Slovenia, and his doctorate in bioinformatics from the University of Zurich, Switzerland. His group developed iCLIP, a transcriptome-wide method that identifies protein-RNA and RNA-RNA interactions in living cells with high resolution. In 2013, he moved with his group to the Institute of Neurology at University College London, and since 2016 the group is based in the Francis Crick Institute. Ule received a doctorate in molecular neuroscience from the Rockefeller University, New York, and started his research group at the MRC Laboratory of Molecular Biology in Cambridge. Gregor Rot, a postdoctoral researcher at the Institute of Molecular Life Sciences, at the University of Zurich. Jernej Ule, group leader at the Francis Crick Institute, and Dr. They will also learn why is it helpful to derive an RNA map, and what does it show us about regulatory mechanisms. Through this webinar, sponsored by Lexogen, attendees will learn how to use the expressRNA platform and analyse 3´ mRNA-Seq QuantSeq data. Conclusions were made that TDP 43 directly regulates diverse types of pre-mRNA processing events according to common positional principles. The team used this approach to show that TDP-43, an RBP involved in several neurodegenerative diseases, binds close to the polyA site to repress, and further downstream to enhance their use. The RNAmotifs2 software also identifies clustered sequence motifs that mediate the regulation of these sites. This reveals at nucleotide resolution the ‘RNA maps’, which demonstrate that RBPs bind to specific positions on pre-mRNAs to regulate the polyA sites. 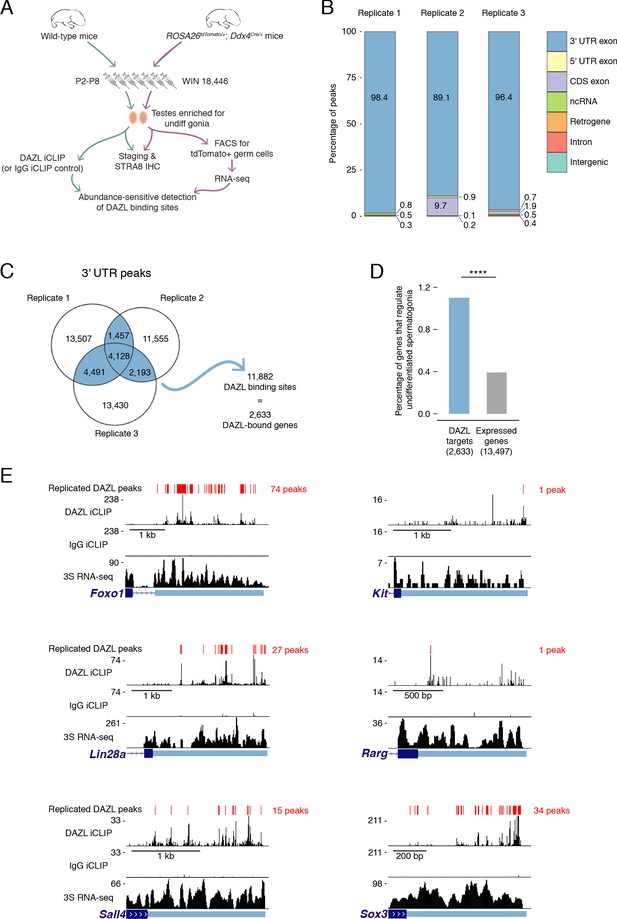
To understand their regulatory principles, the research team developed expressRNA, a web platform encompassing computational tools for integration of the QuantSeq 3´ mRNA-Seq, iCLIP and RNA motif analyses. 
Many RNA binding proteins (RBPs) regulate the selection of alternative polyA sites.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
